Our Research
Wiring the fly brain through genetically hardwired mechanisms
The fly visual system is particularly well suited to studying hard wired mechanisms of brain wiring. There are several reasons for this. First, there is a dense nearly complete map of all cell types within the visual system (a couple hundred) and synaptic connections between them. Second, the patterns of gene expression for most cell types has been determined during development and in the adult. And third, there is vast array of genetic tools available to manipulate different neuron types to explore how molecules regulate specific connections between neurons.
Our studies have led to the discovery of the roles of many cell surface recognition molecules that allow axons and dendrites of developing neurons to discriminate between potential synaptic partners. Neural activity, prior to vision and thus independent of neural experience also contributes to the assembly of neural circuitry. These studies, others in C.elegans and Drosophila, as well as in the developing vertebrate brain have begun to uncover the logic by which the specificity of synaptic connectivity is established during development.
Current studies in the lab are focused on how cell recognition molecules select synaptic partners, form synapses in only very restricted regions of axons and dendrites, how neural activity regulates gene expression during synapse formation, and dissecting the transcriptional strategies and mechanisms regulating wiring programs in different types of neurons.
Uncovering how experience regulates brain wiring in the mouse
Discoveries made about half century ago established that postnatal experience plays an important role in regulating brain wiring in mammals. The periods during postnatal development where the brain is particular sensitive to experience are called critical periods. Different functions have different critical periods with maturation of sensory systems occurring early in postnatal development and higher cognitive functions later.
The role of experience has been particularly well studied in primary visual cortex in the context of vision. We have explored this question in the mouse visual cortex through live imaging studies of the response of neurons to different visual stimuli and the role of vision in instructing this process. These studies have been pursued with the labs of Josh Trachtenberg and Dario Ringach, our colleagues at UCLA. In parallel studies we have been studying how vision regulates the development of cells within the primary visual cortex using single cell transcriptomics, mRNA localization studies and genetics. These studies have been carried out in collaboration with Karthik Shekhar’s lab at University of California, Berkeley. Our studies led to the discovery that molecular specification of cell types requires vision and that this specification parallels the vision-dependent maturation of the functional specification of these neurons.
Our current studies are focused on understanding the relationship between specific cell types in the cortex, how visual experience influences the development of different cell types and cellular processes in the developing postnatal brain, how cell recognition molecules influence synapse maturation, and how these changes are related to vision-dependent ethologically relevant behaviors.
Our Work In Images
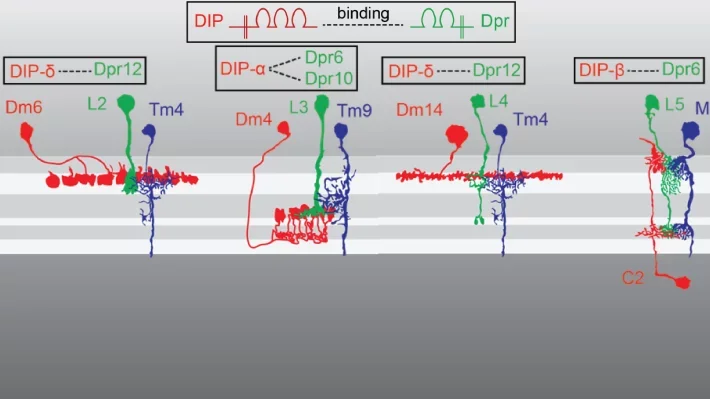
Our recent work has led to the identification of Dpr/DIP protein families as potential regulators of synaptic specificity between neurons in Drosophila. Different Dpr/DIP pairs are seen on different synaptic partners.
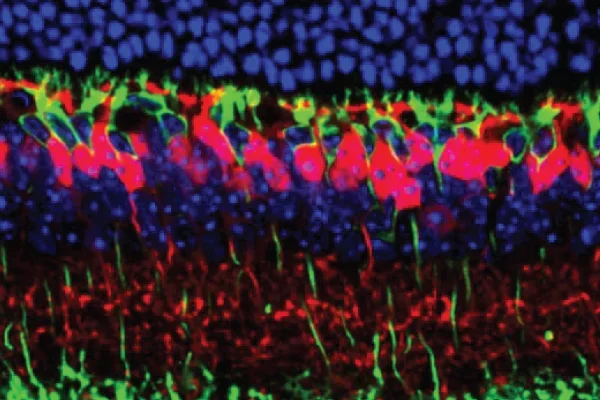
We have used RNA sequencing technology to discover molecular pathways regulating the patterning of the mouse retina. We are exploring how dendrites of rod bipolar neurons (green) and cone bipolar neurons (red) form connections with rod and cone photoreceptors.
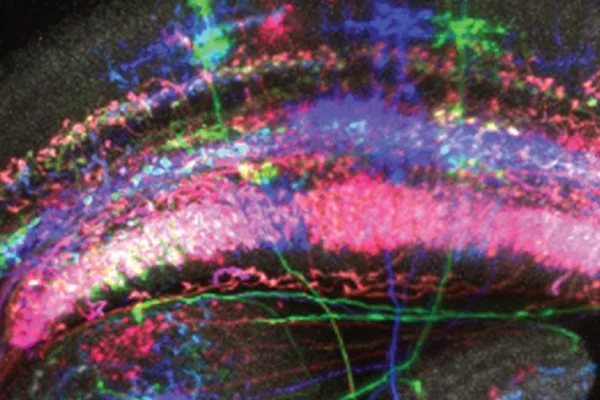
Using the multi-cell-flip-out (MCFO) technology we can identify all the cell types expressing a cell recognition molecule called DIP-epsilon in the medulla region of the fly visual system.